
Genes | Free Full-Text | Transferability and Polymorphism of SSR Markers Located in Flavonoid Pathway Genes in Fragaria and Rubus Species

Development of Genome-wide SSR Markers for Physical Map Construction with PCR-based Polymorphic SSRs in Jute (Corchorus Spp.) | SpringerLink
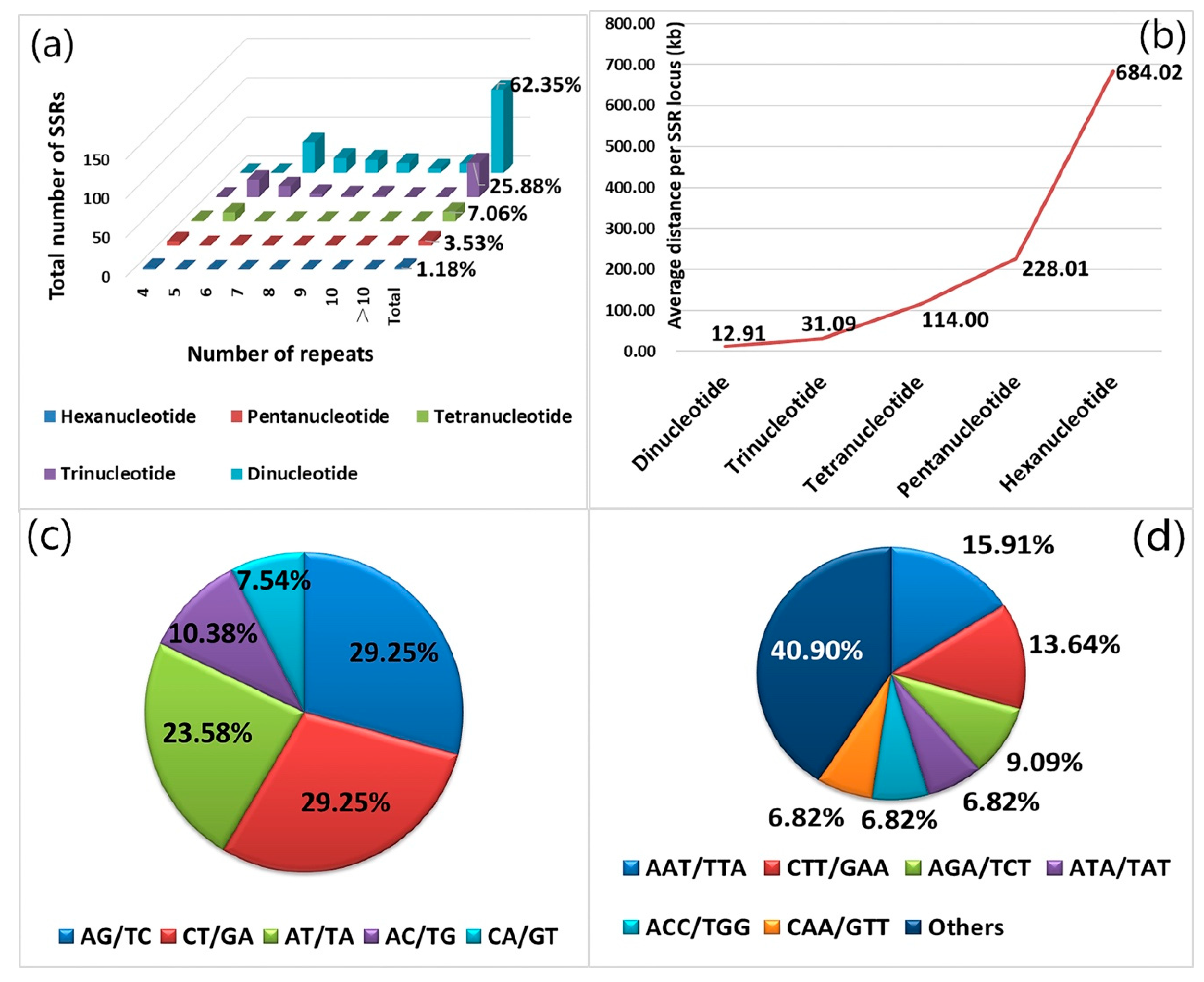
Forests | Free Full-Text | Development and Application of EST-SSR Markers for DNA Fingerprinting and Genetic Diversity Analysis of the Main Cultivars of Black Locust (Robinia pseudoacacia L.) in China
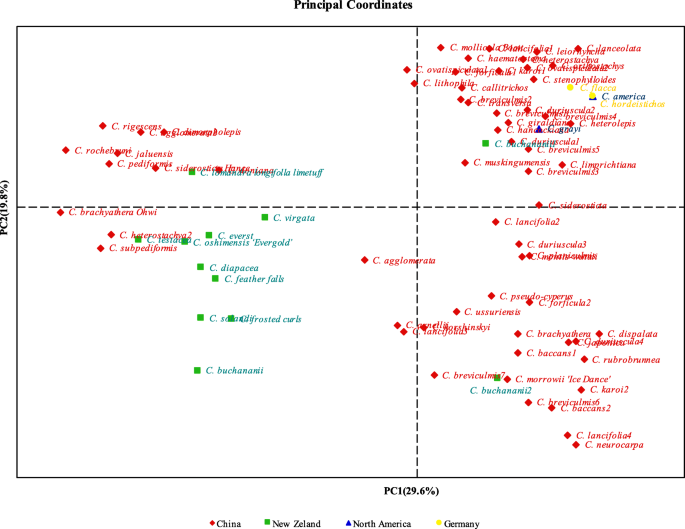
The development of SSR markers based on RNA-sequencing and its validation between and within Carex L. species | BMC Plant Biology | Full Text
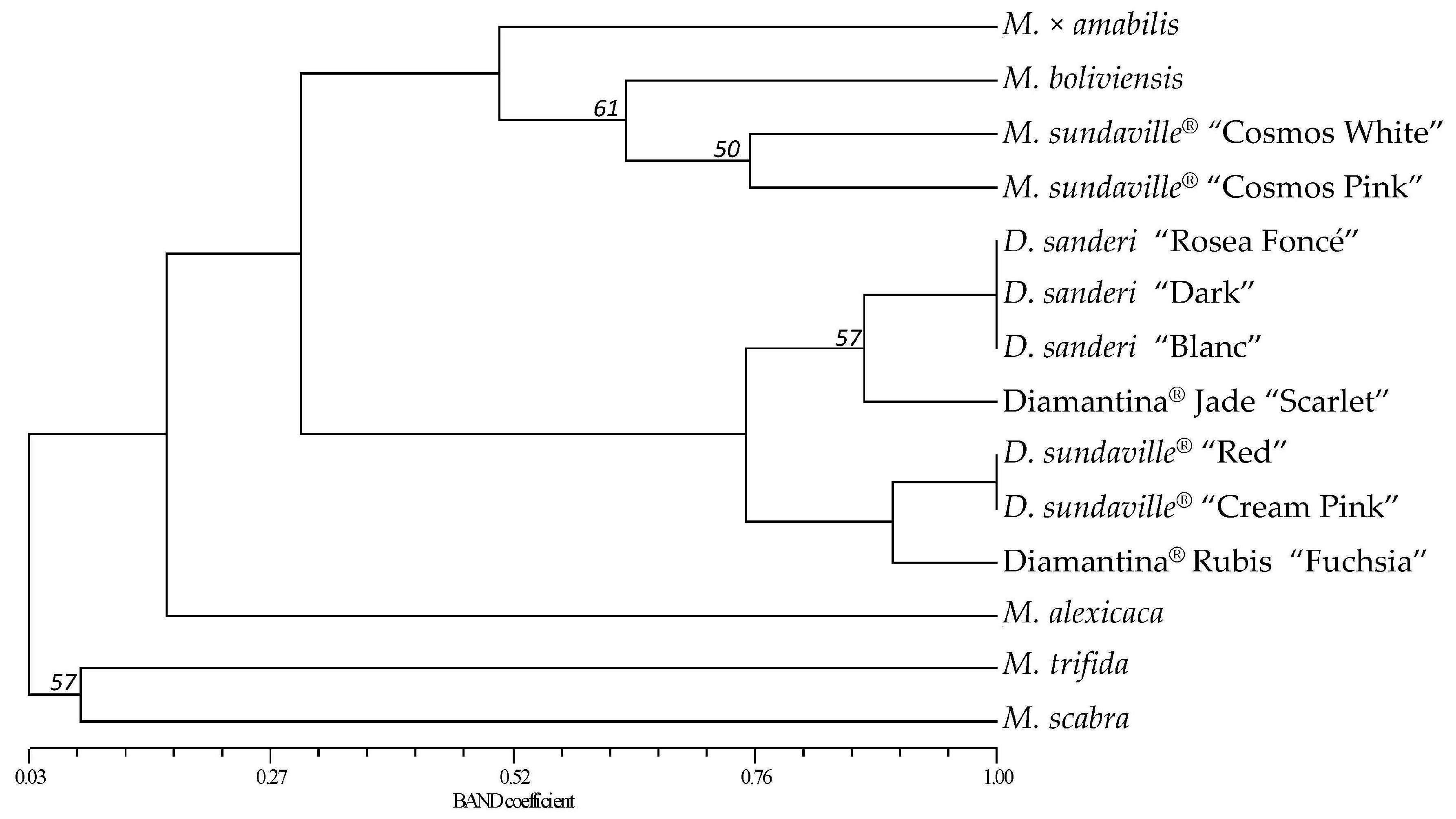
Molecules | Free Full-Text | A Set of 20 New SSR Markers Developed and Evaluated in Mandevilla Lindl.

Polymorphism detected by ten SSR markers among 18 genotypes of three... | Download Scientific Diagram
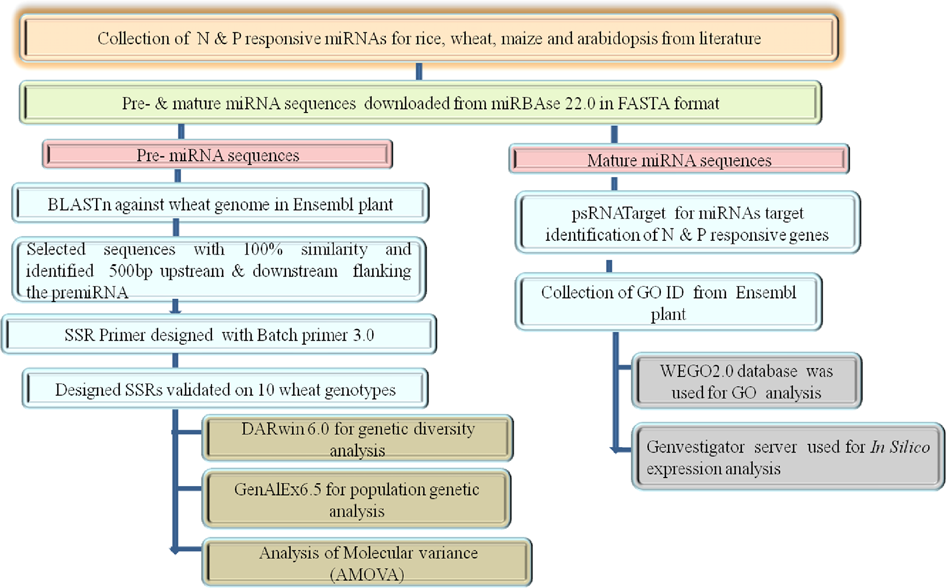
Development and characterization of nitrogen and phosphorus use efficiency responsive genic and miRNA derived SSR markers in wheat | Heredity
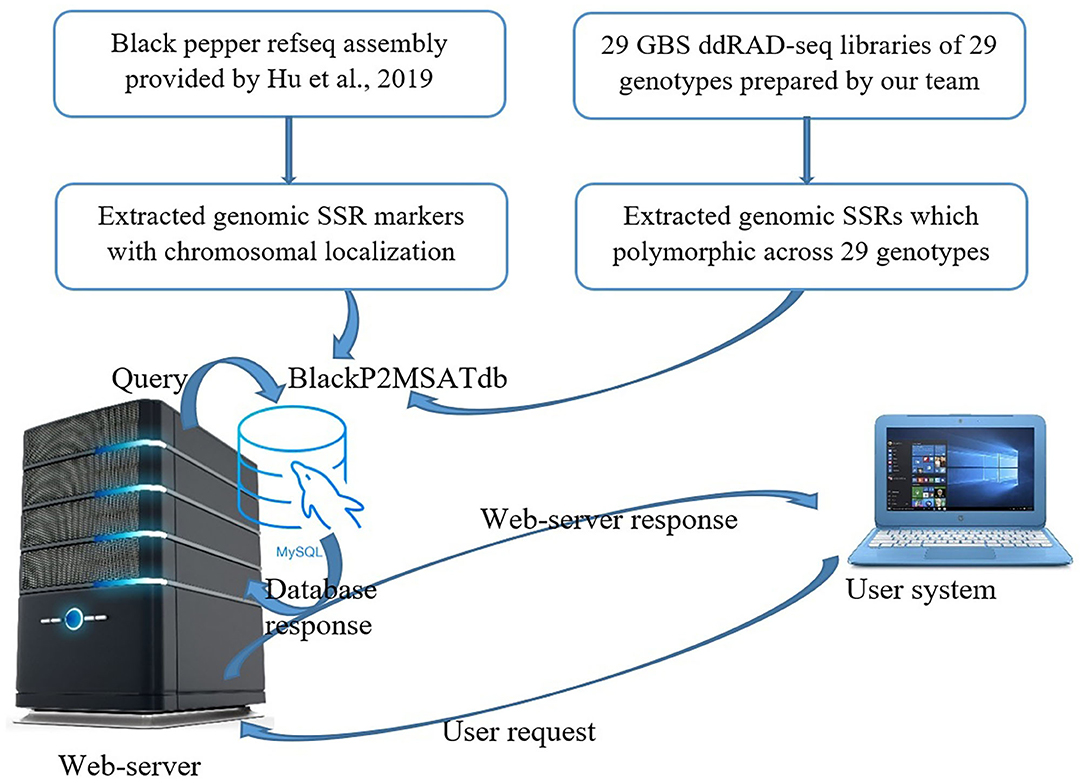
Frontiers | Rapid Genome-Wide Location-Specific Polymorphic SSR Marker Discovery in Black Pepper by GBS Approach
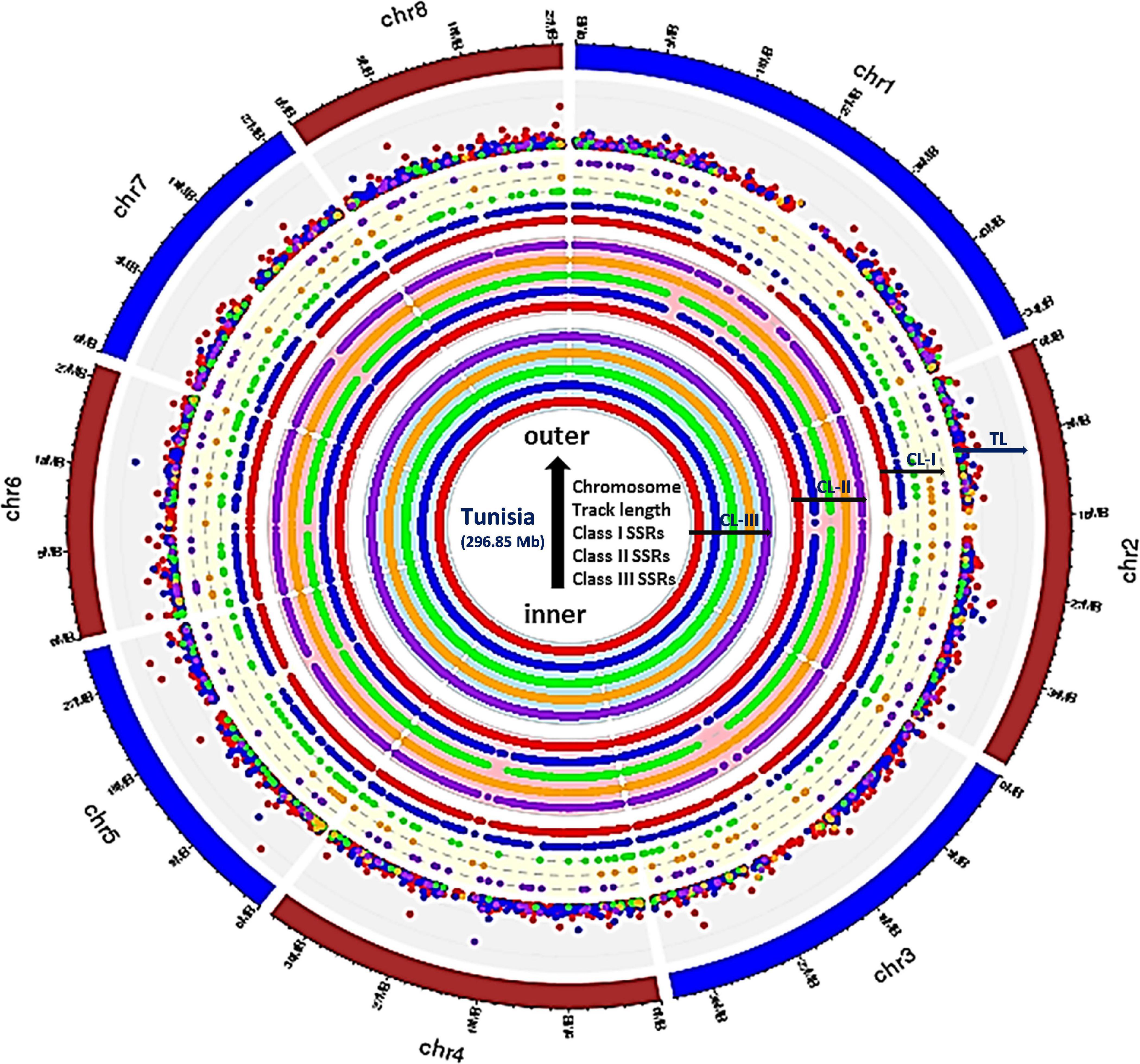
Frontiers | Comprehensive Characterization and Validation of Chromosome-Specific Highly Polymorphic SSR Markers From Pomegranate (Punica granatum L.) cv. Tunisia Genome

Characterization and Development of EST-SSR Markers from Transcriptome Sequences of Chrysanthemum (Chrysanthemum ×morifolium Ramat.) in: HortScience Volume 54 Issue 5 (2019)

MultiplexSSR: A pipeline for developing multiplex SSR‐PCR assays from resequencing data - Guo - 2020 - Ecology and Evolution - Wiley Online Library
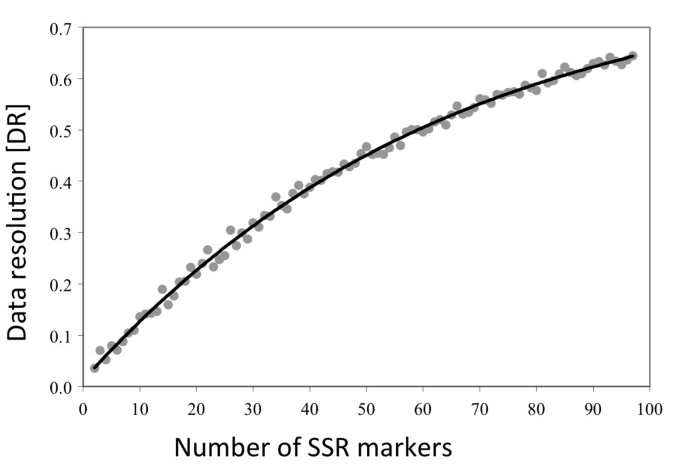
Development of genomic SSR markers for fingerprinting lettuce (Lactuca sativaL.) cultivars and mapping genes | BMC Plant Biology | Full Text
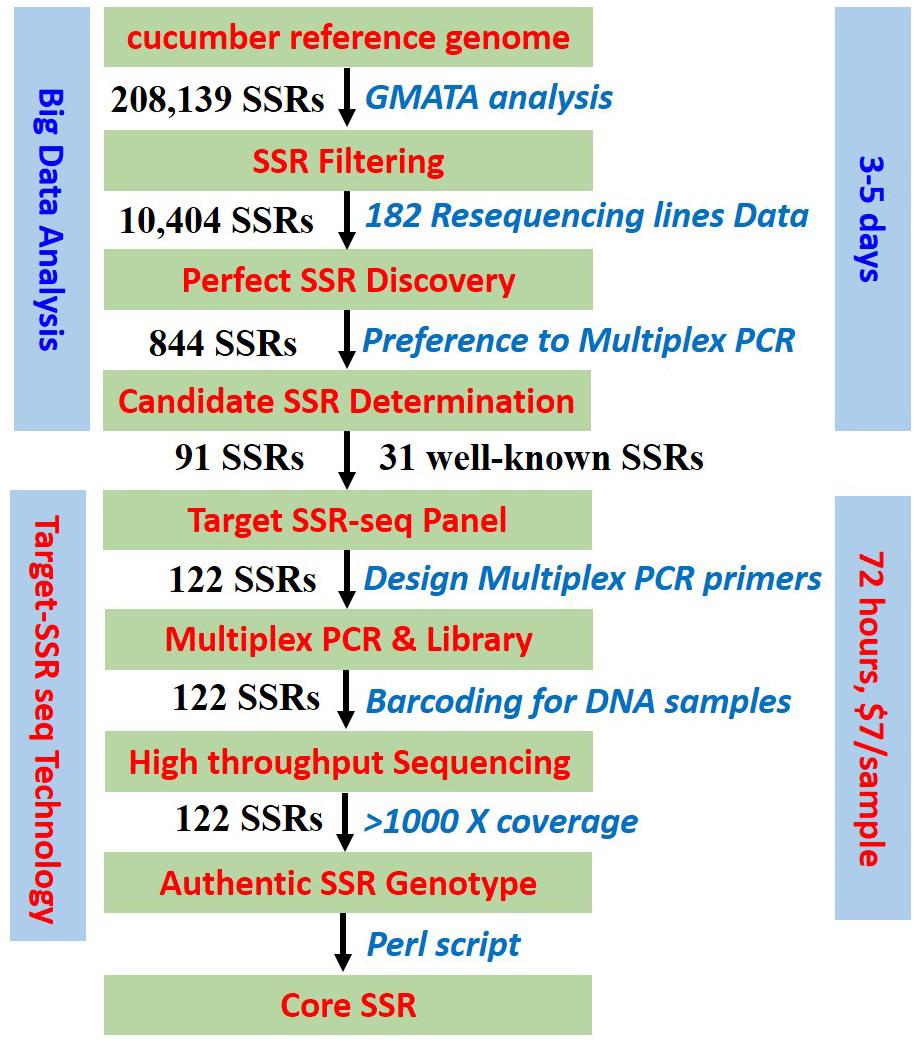
Frontiers | Target SSR-Seq: A Novel SSR Genotyping Technology Associate With Perfect SSRs in Genetic Analysis of Cucumber Varieties

Cross-species transferability of genomic SSR markers and genetic diversity among Asparagus racemosus Willd. Accessions - ScienceDirect
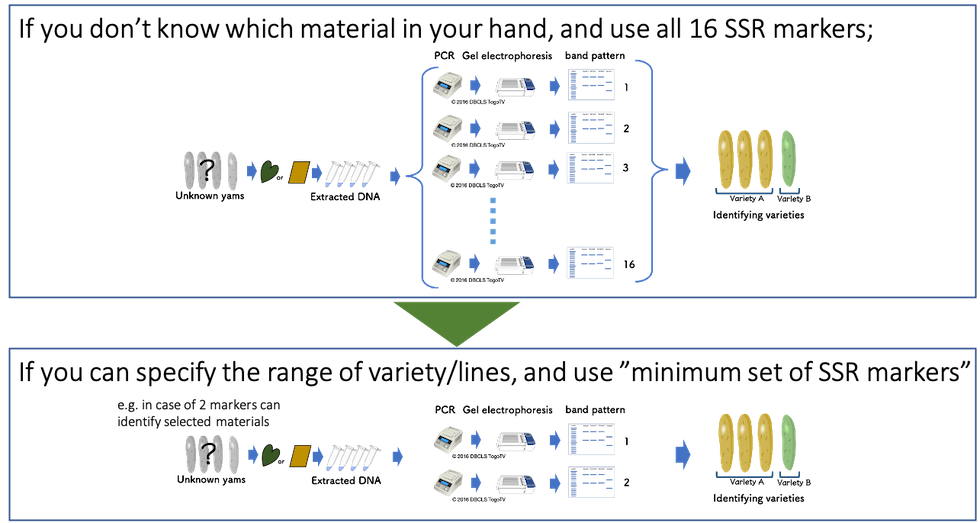
Yam Variety Identification Toolkit:Step 1 | Japan International Research Center for Agricultural Sciences | JIRCAS
![Frontiers | New Hypervariable SSR Markers for Diversity Analysis, Hybrid Purity Testing and Trait Mapping in Pigeonpea [Cajanus cajan (L.) Millspaugh] Frontiers | New Hypervariable SSR Markers for Diversity Analysis, Hybrid Purity Testing and Trait Mapping in Pigeonpea [Cajanus cajan (L.) Millspaugh]](https://www.frontiersin.org/files/Articles/243873/fpls-08-00377-HTML-r1/image_m/fpls-08-00377-g001.jpg)
Frontiers | New Hypervariable SSR Markers for Diversity Analysis, Hybrid Purity Testing and Trait Mapping in Pigeonpea [Cajanus cajan (L.) Millspaugh]

Genome-wide identification of simple sequence repeats and development of polymorphic SSR markers in swamp eel (Monopterus albus) - Hai-feng Tian, Qiao-mu Hu, Zhong Li, 2021
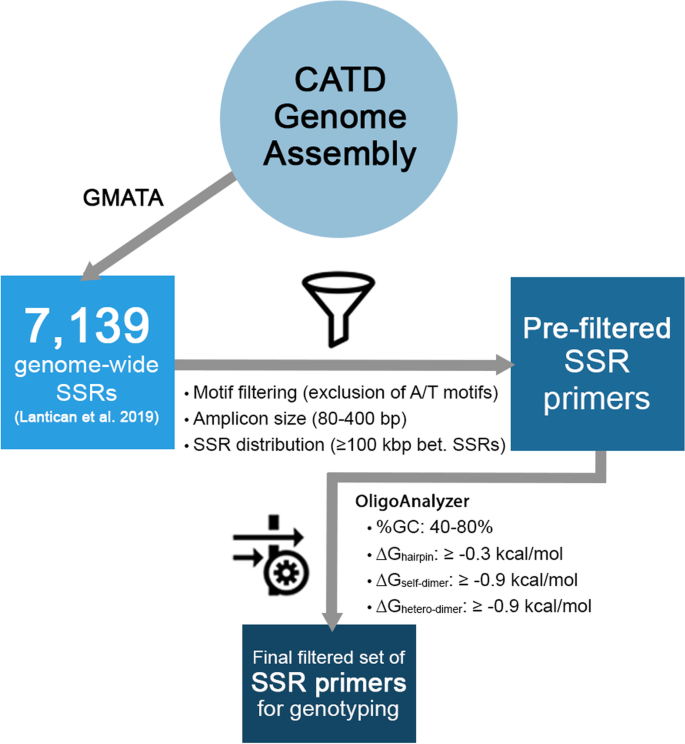
Mining and validation of novel simple sequence repeat (SSR) markers derived from coconut (Cocos nucifera L.) genome assembly | Journal of Genetic Engineering and Biotechnology | Full Text
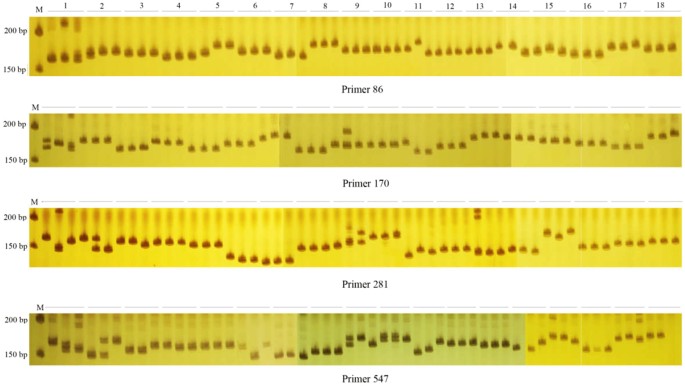

Cross-species transferability of EST-SSR markers developed from the transcriptome of Melilotus and their application to population genetics research | Scientific Reports






